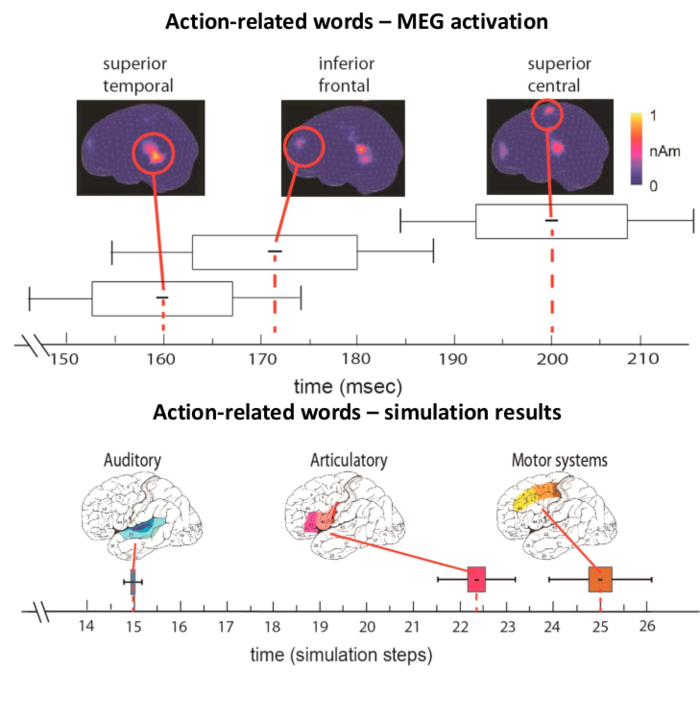
Comparing Human and Modelled Brains
Comparing maximal activation latencies in human brains and simulated brain networks.
Image Credit: Adapted from Tomasello et al., 2017.
In the Experimental Research section, we have described our approaches to data collection for comparison with the computational model. But how can we actually compare data from sources as different as computer simulations, human brains and macaque or chimpanzee brains? On top of that, how can we compare the results from fMRI and EEG experiments and relate them to the stimuli presented in the experiments?
When we compare model and real-life data, or even data from two different measurement methods, we are looking to answer one question: how similar are the two? For example, EEG experiments yield a measure of postsynaptic electric potentials and fMRI experiments yield a measure of blood-oxygen levels in the brain as different signatures of brain activity. One option is to use these brain measures and calculate their effect on results obtained from neural network simulations, for example neural firing rates over time. However, this strategy may be based on weak assumptions and uncertainties. An alternative way is to directly compare the different signatures in relation to the stimuli which caused them.
Instead of directly comparing voltages and oxygen concentration with neural network outputs, we compare the similarity structure of each measure. Stimuli themselves have different levels of similarity, brain activations also sometimes resemble each other more or less, and likewise, the networks’ activities may be similar to each other or different. The degree of similarity can be determined using precise mathematical measures. We here use a method proposed by Nikolaus Kriegeskorte and colleagues (Kriegeskorte et al., 2008): for each stimulus pair, their similarity can be mapped in a similarity matrix, and likewise, the similarity level of the brain activation patterns elicited by these stimuli can be calculated. Finally, analogous calculations of similarities can be done for neural network simulations. Correlation between different domains (stimulus, brain activity, network activity) can be calculated to determine their match or mismatch. In this way, data from different sources (e.g. stimuli, real brains, simulations) can be compared by looking for their similarity structure, i.e. the degree of match between their respective similarity matrices.
By thus abstracting away from specific measures, we can compare across data origins (human – computer model – chimpanzee), experimental method (fMRI – EEG) and even across different brain areas (primary visual cortex – primary auditory cortex). In research on semantics, we can also use this technique to determine how similar the representations of different concepts – for example ‘apples’ and ‘pears’ – are.
For an example of the method in action, consider the following two papers: Francesca Carota and colleagues use it to investigate whether the semantic knowledge in the brain is organized more like a taxonomic knowledge base (a bird is a type of animal) or according to principles of distributional semantics (the words bird and fly often cooccur) (Carota et al., 2021), while Malte Henningsen-Schomers and colleagues use RSA on the output of a brain-constrained neural network to investigate how similarly it represents different learned concepts – think of ‘apples’ vs ‘pears’ again (Henningsen-Schomers et al., under review).
References
Carota, F., Nili, H., Pulvermüller, F., & Kriegeskorte, N. (2021). Distinct fronto-temporal substrates of distributional and taxonomic similarity among words: Evidence from RSA of BOLD signals. NeuroImage, 224, 117408. https://doi.org/10.1016/j.neuroimage.2020.117408
Henningsen-Schomers, M. R., Garagnani, M., & Pulvermüller, F. (under review). Influence of language on concept formation and perception in a brain-constrained deep neural network model.
Kriegeskorte, N., Mur, M., & Bandettini, P. (2008). Representational similarity analysis - connecting the branches of systems neuroscience. Frontiers in Systems Neuroscience, 2, 4. https://doi.org/10.3389/neuro.06.004.2008